Black Cohosh Adulteration Laboratory Guidance Document
By Stefan Gafner, PhD
Chief Science Officer, American Botanical Council
Technical Director, ABC-AHP-NCNPR Botanical Adulterants Program
Keywords: Adulterant,
adulteration, black cohosh, Actaea
cimicifuga, Actaea dahurica, Actaea heracleifolia, Actaea racemosa, Chinese cimicifuga, Cimicifuga racemosa
CONTENTS
1. Purpose
2. Scope
3. Common and scientific names
3.1 Common Name
3.2 Other Common Names
3.3
Latin Binomial
3.4 Synonyms
3.5 Botanical Family
4. Botanical Description
Table 1. Scientific names, family, and common names
of known black cohosh adulterants
5. Identification and Distinction using
Macroanatomical Characteristics
6. Identification and Distinction using
Microanatomical Characteristics
7. Genetic Identification and Distinction
8. Chemical Identification and
Distinction
8.1 Chemistry of Actaea racemosa and the Potential Adulterants
Figure 1: Major triterpene glycosides in
black cohosh
Figure
2: Phenolic acids found in black cohosh
Table
2. Nomenclature of major triterpene glycosides from A. racemosa according to Qiu et al.
Figure
3: Structures of cimifugin (15) and cimifugin-3-O-glucoside (16)
8.2
Laboratory Methods
8.2.1
HPTLC
Figure 4: HPTLC analysis of root extracts of black
cohosh, Actaea podocarpa (syn. Cimicifuga americana), and two Chinese Actaea spp., A. dahurica (syn. C. dahurica)
and A. cimicifuga (syn. C. foetida). Detection: visible light
after derivatization with sulfuric acid reagent. Image provided by CAMAG
(Muttenz, Switzerland).
Figure 5: HPTLC evaluation of black cohosh
adulteration with Actaea cimicifuga
and A. simplex. Detection: UV light
(366 nm) after derivatization with boric acid/oxalic acid reagent. Lane 1:
cimifugin, actein, isoferulic acid (with increasing Rf value); lane 2: A. racemosa; lane 3: 5% A. cimicifuga in A. racemosa; lane 4: 5% A.
simplex in A. racemosa. Image
provided by CAMAG (Muttenz, Switzerland).
8.2.2
HPLC and UHPLC
8.2.3
MS-Fingerprinting
8.2.4
NMR
9. Conclusion
Table 3. Comparison among the different
approaches to authenticate A. racemosa.
10. References
Appendix 1
Table 4: Comments on the published HPLC methods for A. racemosa.
1. Purpose
In recent years, adulteration of black cohosh roots and rhizomes (Actaea racemosa, Ranunculaceae) has become more apparent.
Adulteration predominantly occurs with Chinese species of Actaea such as A.
heracleifolia, A. dahurica, and A. cimicifuga (all known by the common
name Chinese cimicifuga and by the Chinese name of sheng ma). Additionally, the Chinese cimicifuga (sheng ma) market is commonly adulterated
with Serratula chinensis (guang dong sheng ma [Asteraceae]). Adulteration has also been reported with North American Actaea species growing in the same area
as black cohosh, such as A. pachypoda, A. rubra, and A. podocarpa. This Laboratory
Guidance Document presents a review of the various analytical technologies used
to differentiate between authentic A. racemosa and its potentially adulterating species.
2. Scope
The various analytical
methods were reviewed with the specific purpose of identifying strengths and
limitations of the existing methods for differentiating A. racemosa from its potentially adulterating species. Analysts can
use this review to help guide the appropriate choice of techniques for their
specific black cohosh products for qualitative purposes. The recommendation of
a specific method for testing A. racemosa
materials in their particular matrix in
this Laboratory Guidance Document does not reduce or remove the responsibility
of laboratory personnel to demonstrate adequate method performance in their own
laboratories using accepted protocols outlined in the United States Food and
Drug Administration’s Good Manufacturing Practices (GMPs) rule (21 CFR Part 111)
and those published by AOAC International, International Organization for
Standardization (ISO), World Health Organization (WHO), and International
Conference on Harmonisation (ICH).
3.
Common and scientific names
3.1 Common Name: Black cohosh
Note:
According to the American Herbal Products Association’s Herbs of Commerce, 2nd ed.,1 the standard common name
of Actaea racemosa is black cohosh.
Any other species referred to as black cohosh is considered an adulterant under
botanical dietary supplement GMPs. Each of the other Actaea species has their own common name.
3.2 Other Common Names
English: Black bugbane, snakeroot, rheumatism weed, fairy candle, tall bugbane,
macrotys, macrotrys, battleweed, columbine-leaved leontice, cordate rattle top,
rattleweed, false cohosh, papoose root
Chinese: Zong zhuang sheng ma (总状升麻)
French: Actée à grappes
noires, cimicaire à grappes, chasse-punaises, cimifuge, herbe à punaise,
serpentaire noire
German:
Traubensilberkerze, Wanzenkraut, Frauenwurzel, langtraubiges Christophskraut,
Nordamerikanische Schlangenwurzel, schwarze Schlangenwurzel
Italian:
Cimicifuga, serpentaria nera, radice della squaw, actea nera
Spanish: Cohosh negro, raíz de culebra negra
3.3 Latin Binomial: Actaea racemosa L.
3.4 Synonyms: Cimicifuga racemosa
(L.) Nutt.
3.5 Botanical Family:
Ranunculaceae
4.
Botanical Description
Actaea
racemosa is native only to the eastern portion of North America,
although it is propagated in numerous countries. Botanical descriptions for A. racemosa and its adulterant species
are provided in local, national, and international floras, including the Flora of North America and the Flora of China (for Chinese species of Actaea and Vernonia). Additionally, a botanical description for Actaea species reported as adulterants
is provided in the black cohosh monograph of the American Herbal Pharmacopoeia
(AHP) and is accompanied by illustrations and images.2 A detailed
morphological analysis of black cohosh populations was published by Gardner et
al.3 Identifying and differentiating among the species requires
personnel trained in botany and requires the assessment of materials whose
botanical characteristic features are intact.
Table 1.
Scientific names, family, and common
names of known black cohosh adulterants*
|
Speciesa
|
Synonym(s)b
|
Family
|
Common namec
|
Other common namesd
|
|
Actaea
cordifolia DC.
|
A.
rubifolia (Kearney) Kartesz;
Cimicifuga
rubifolia Kearney;
C. racemosa var. cordifolia (DC.) A. Gray
|
Ranunculaceae
|
|
Appalachian
bugbane
|
|
Actaea pachypoda Elliott
|
A. brachypetala
var. coerulea DC.;
A. brachypetala
var. microcarpa DC.;
A. pachypoda
f. microcarpa (DC.) Fassett
|
Ranunculaceae
|
|
White
baneberry, doll’s eyes, necklace weed
|
|
Actaea podocarpa
DC.
|
A. americana Prantl;
C. americana Michx.;
C. podocarpa
(DC.) Elliott
|
Ranunculaceae
|
|
Yellow
cohosh, mountain bugbane
|
|
Actaea rubra (Aiton) Willd.
|
A. arguta
Nutt.
A. caudata
Greene;
A. erythrocarpa (Fisch.) Kom.;
A. rubra subsp. arguta (Nutt.) Hultén;
A. viridiflora
Greene
|
Ranunculaceae
|
|
Red
baneberry
|
|
Actaea cimicifuga L.
|
C. foetida L.
|
Ranunculaceae
|
Chinese cimicifuga
|
Skunk
bugbane,
xi sheng ma (西升麻)e, sheng ma (升麻)f
|
|
Actaea dahurica (Turcz. ex Fisch. & C.A. Mey.)
Franch.
|
A. pterosperma Turcz. ex Fisch. & C.A. Mey.;
Actinospora
dahurica Turcz. ex Fisch. & C.A. Mey.;
C. dahurica (Turcz.) Maxim.
|
Ranunculaceae
|
Chinese
cimicifuga
|
Dahurian
bugbane,
bei sheng ma (北升麻)e, xing an sheng
ma (兴安升麻)f
|
|
Actaea heracleifolia (Kom.) J.Compton
|
C. heracleifolia Kom.
|
Ranunculaceae
|
Chinese
cimicifuga
|
Large-leaf
bugbane, guan sheng ma (关升麻)e,
da san ye sheng ma(大三叶升麻)f
|
|
Actaea simplex (DC.) Wormsk. ex Prantl
|
A. cimicifuga var. simplex DC.;
C. simplex (DC.) Wormsk. ex Turcz. ;
C. ussuriensis
Oett., Thalictrodes simplex (DC.) Kuntze
|
Ranunculaceae
|
|
dan
sui sheng ma (单穗升麻)f
|
|
Acilepis aspera (Buch.-Ham.) H.Rob.
|
Vernonia aspera Buch.-Ham.;
V.
roxburghii Less.;
V.
teres Wall.;
Xipholepis aspera (Buch.-Ham.) Steetz
|
Asteraceae
|
|
cao ye ban jiu ju (糙叶斑鸠菊)f
|
|
Serratula chinensis S. Moore
|
Centaurea
missionis H. Lév.
|
Asteraceae
|
|
guang dong sheng ma
(
广东升麻)e,
hua ma hua tou (华麻花头)f
|
*Plants listed in this table
include plants that have historically been known to be adulterants of black
cohosh but do not necessarily reflect plants that are known to be of
significant potential or actual adulteration in the herb marketplace in recent
years.
aThe Plant
List and the Tropicos database.4,5
bThe Plant
List and the Tropicos database.5 A comprehensive list of synonyms
can be accessed through both websites.
cHerbs of Commerce, 2nd
ed.1
dHerbs of Commerce, 2nd
ed.,1 the USDA PLANTS Database,6 and Pengelly and Bennett.7
eChinese common name
in trade.
fChinese scientific botanical name.
Note:
For Actaea/Cimicifuga, the accepted species
name of the species may differ between The Plant List and the Tropicos database;
The Plant List lists Actaea cimicifuga
L. as the accepted Latin binomial, while Tropicos lists Cimicifuga foetida L. In such cases, the names from The Plant List
are indicated in this table as the accepted species name.
It is believed that Chinese
material belonging to the Actaea or Cimicifuga genera other than the species
identified above is sometimes sold as “black cohosh.”8 Hagers Handbuch der Drogen und Arzneistoffe
also lists other Actaea species, and
the roots of arnica (Arnica montana,
Asteraceae) and the rhizomes of black hellebore (Helleborus niger, Ranunculaceae) as possible adulterants.9
However, there is no evidence that adulteration with arnica or black hellebore
is still an issue in the current marketplace.
Sections 5-8 of this
document discuss macroscopic, microscopic, genetic, and phytochemical
authentication methods for A. racemosa.
A comparison among the various approaches is presented in Table 3 at the end of
section 9.
5.
Identification and Distinction using Macroanatomical Characteristics
Macroscopic identification
criteria for A. racemosa have been
published in the AHP monograph,2 in the book chapter by Hiller,10
and in the US Pharmacopeia (USP).11 However, there is no information
on criteria to distinguish black cohosh macroscopically from related Actaea species and such a distinction
may be challenging. Therefore, other means for authentication should be used.
The AHP monograph does have photographs of authentic A. racemosa and A. podocarpa,
as well as Chinese samples. These species are morphologically distinguishable
from each other.
6.
Identification and Distinction using Microanatomical Characteristics
Detailed
microscopic descriptions of A. racemosa
are found in numerous references.2,12-14 In addition, the Botanical Microscopy Atlas12
contains images of transverse sections of the root and rhizome of A. pachypoda and A. podocarpa. Applequist lists the microscopic
characteristics of North American species of Actaea growing in the same area. Whole and cut root material of A. racemosa, A. pachypoda, and A.
podocarpa can be distinguished based on the number and shape of xylem
strands in the rhizome and the root. Actaea
rubra is less easily distinguished from A.
racemosa, and, even if there are differences in the number and width of the
vascular bundles, experience and much attention to detail is required to distinguish
the two species.14 The AHP monograph
contains illustrations and photographic images of authentic A. racemosa, but not its adulterants. In
the Chinese literature, there is information related to the microscopic
differentiation of a number of Actaea
species, including the three official species listed by the Chinese Pharmacopoeia for sheng ma, a number of other species
from the same genus, and common adulterants.15 The characteristic
features for both transverse sections and powder forms are covered. Generally
speaking, the differences among the Chinese Actaea
species are subtle and it is challenging to differentiate them because they
share many similar microscopic features. However, a comparison of the
microscopic features of Chinese Actaea
materials with black cohosh is lacking.
Based on the available
authoritative resources, there is no
single reference that contains information on A. racemosa and all its known current adulterants, e.g., the
Chinese Actaea species, co-occurring
Appalachian species, Acilepis aspera, or Serratula
chinensis.
Comments:
While microscopic distinction of A.
racemosa and closely related North American Actaea species using transverse sections of whole roots has been
described, it is unclear if a distinction can be achieved with powdered root
material. To the best of the knowledge of the author and peer reviewers of this
document, there is no detailed comparison of microanatomical characteristics
among black cohosh and Chinese Actaea species.
Therefore, the sole use of microscopy for the authentication of A. racemosa and for the detection of its
adulterants should be considered inadequate.
7.
Genetic Identification and Distinction
Methods described in the
following literature were evaluated in this review: Zerega et al.16
and Baker et al.17
Comments: Several
scientists have developed DNA-based methods for Actaea species identification and detection of adulterants. In 2002,
Zerega et al.16 published a method using Amplified Fragment Length Polymorphism
(AFLP); however, newer methods utilizing DNA sequence-based methods are
superior in specificity and reproducibility. In the publication by Baker et al.,17
DNA barcodes that amplify specific regions of the genome that are variable among
Actaea species are used. DNA
sequence-based methods utilizing “universal” (vs. species-specific) primers can
also amplify unexpected adulterants and can provide an extremely reliable and
robust system not only for distinguishing among closely related Actaea species, but also in detecting
mixtures with other adulterants. In a collaboration between industry and the
National Institute of Standards and Technology (NIST), A. racemosa leaf and root Standard Reference Materials (SRMs) with
associated DNA barcodes for two validated gene regions have been developed;
these DNA barcodes were validated for specificity across numerous samples of
target and adulterant species and have demonstrated a 100% probability of
identification.18 The NIST SRMs with associated DNA barcodes and
validation data will be made available by the end of 2015 (Catherine Rimmer
e-mail communication, June 25, 2015). This authentication method has been
successfully utilized across a wide range of starting materials, including
dried and fresh leaves and roots in whole, cut, and powdered form; however,
validated methods for more highly processed materials such as dried extracts
are not currently available. Because genetic identification is unable to
determine plant part, morphological, microscopic, or chemical evaluation is
also necessary for materials in powdered form.
8.
Chemical Identification and Distinction
A large number of analytical
methods have been published that are used for identifying A. racemosa roots/rhizomes and root/rhizome extracts based on their
chemistry. These methods are cited in the Laboratory Methods section below. For
some methods, distinction based on the phytochemical profile may require a
detailed knowledge of the constituents of black cohosh and its adulterants. The
important components in A. racemosa and
its adulterating species are listed below. When distinction is based on
chromatographic or spectral patterns, identification of specific constituents
may not be necessary.
8.1
Chemistry of Actaea racemosa and the Potential
Adulterants
Actaea racemosa:
Besides the prominent triterpene glycosides (Figure 1) of the 9,19-cycloartenol
type, phenolic acids, tannins, fatty acids, and nitrogen-containing compounds
like alkaloids, nucleobases, nucleosides, and phenolic amides and amines have
been described from black cohosh.2,7,10,19-24 Ganzera et al.25
and Avula et al.26,27 identified the major triterpene glycosides as
actein (1), 23-epi-26-deoxyactein (2), and cimiracemoside A (3) (different from cimiracemoside A
isolated by Shao et al.28). Additional triterpene glycosides
occurring in large concentrations are cimicifugoside H-1 (4), cimigenol-3-O-xyloside
(5), cimigenol-3-O-arabinoside (6), 23-O-acetylshengmanol-3-O-xyloside (7), and 23-O-acetylshengmanol-3-O-arabinoside (8).29-33
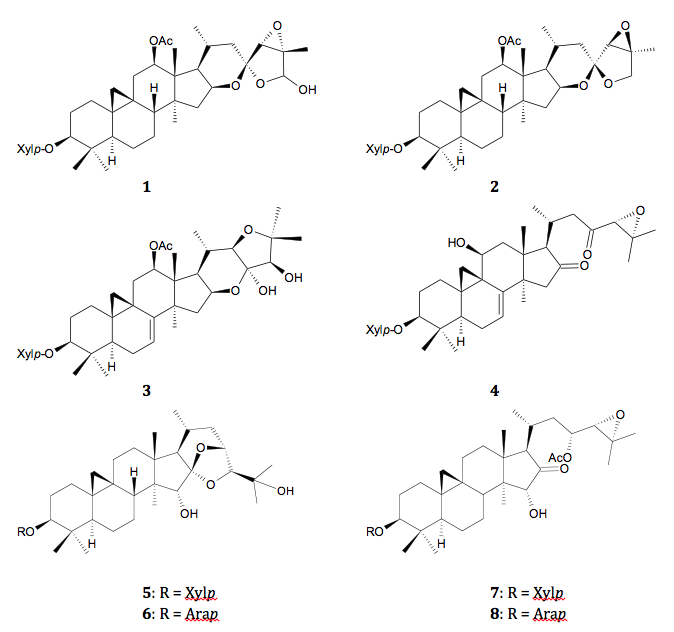
Figure
1: Major triterpene glycosides in black cohosh
The phenolic acids (Figure 2) isolated from black cohosh
are caffeic acid (9), ferulic acid (10), and isoferulic acid (11), either in simple form or as esters
of fukiic acid or piscidic acid, e.g., fukinolic acid (12, syn: cimicifugic acid KC), cimicifugic acids A (13, syn: cimicifugic acid KF), B (14, syn: cimicifugic acid KI), E (syn:
cimicifugic acid PF), and F (syn: cimicifugic acid PI).31,34
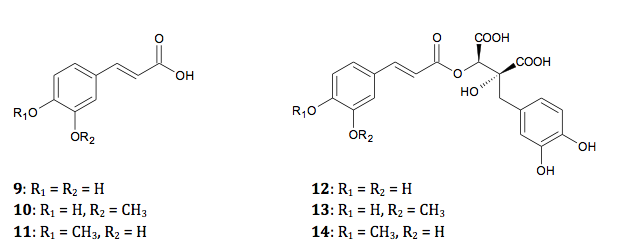
Figure
2: Phenolic acids found in black cohosh
Research to find new phytochemicals
in black cohosh roots and rhizomes has led to a rather well-known phytochemical
profile for this plant, but also to a proliferation of confusing and sometimes
erroneous common names, in particular for the cycloartane triterpenes. A more
rational approach was described by Qiu et al.35; it remains to be
seen if the rather lengthy names proposed in the Qiu et al. paper will be
accepted by the scientific community. Examples of the new names for some of the
major triterpene glycosides are given in Table 2.
Table
2. Nomenclature of major triterpene glycosides from A. racemosa according to Qiu et al.35
|
Original name
|
Synonyms
|
New name
|
|
23-O-acetylshengmanol-3-O-xyloside
(7)
|
|
(23R)-23-acetoxy-(24S)-24,25-epoxy-(15R)-15-hydroxy-16-oxo-3-O-β-d-xylopyranosylactanoside
|
|
Actein (1)
|
Shengmating
|
(12R)-12-acetoxy-(24R,25S)-24,25-epoxy-(26R&S)-26-hydroxy-3-O-β-d-xylopyranosylacta-(16S,23R)-16,23;23,26-binoxoside
|
|
Cimigenol-3-O-arabinoside (6)
|
Cimiracemoside C,
cimicifugoside M
|
(15R)-15,25-dihydroxy-3-O-α-L-arabinopyranosylacta-(16S,23R,24S)-16,23;16,24-binoxoside
|
|
Cimigenol-3-O-xyloside (5)
|
Cimicifugol-xyloside, cimigenoside,
cimicifugoside
|
(15R)-15,25-dihydroxy-3-O-β-d-xylopyranosylacta-(16S,23R,24S)-16,23;16,24-binoxoside
|
|
Cimiracemoside A (3)
|
Cimiracemoside F
|
(12R)-12-acetoxy-7,8-didehydro-(23R,24R)-23,24-dihydroxy-3-O-β-d-xylopyranosylacta-(16S,22R)-16,23;22,25-binoxoside
|
|
23-Epi-26-deoxyactein (2)
|
26-Deoxyactein,
27-deoxyactein
|
(12R)-12-acetoxy-(24R,25R)-24,25-epoxy-3-O-β-d-xylopyranosylacta-(16S,23R)-16,23;23,26-binoxoside
|
Actaea cordifolia: The major phenolic acid in A. cordifolia is fukinolic acid (12). The roots/rhizomes also contain
cimicifugic acids A (13) and B (14). The authors were unable to determine
the identity of the predominant triterpene glycosides, but detected smaller
amounts of 1, 2, 4, and 25-O-acetylcimigenol-3-O-xyloside. In addition, the analysis revealed the occurrence of
the dihydrofurochromone cimifugin (15)
and cimifugin-3-O-glucoside (16) (Figure 3).36 The
analysis of A. cordifolia by a
different group led to the identification of 6 as a major triterpene glycoside, but 1, 2, and 4 were not found.37 Based on
the contradictory results, additional work needs to be done to conclusively
establish the triterpene glycoside composition of A. cordifolia.
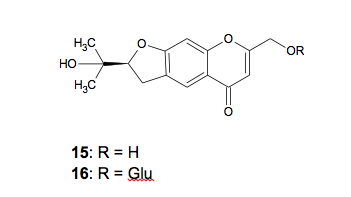
Figure
3: Structures of cimifugin (15) and cimifugin-3-O-glucoside (16)
Actaea pachypoda: The
predominant phenolic acid is 12,
with lesser amounts of 13 and 14.36 Triterpene glycoside
fingerprints of A. pachypoda have
been established by HPLC-MS, and identified 1-3 and 5-7, although
only 1 and 2 were consistently found.26,30,36
Actaea podocarpa: Root/rhizome
material from A. podocarpa was found
to contain mainly 10,
2-feruloylpiscidic acid, and 13, and
only traces of 12 and 14.36,37 The analysis of the
triterpene glycosides showed the presence of 5 and 6. Importantly, 1 was absent and 2 was found in only one of the four samples analyzed.26,30,36,37
A number of unique triterpene glycosides, the podocarpasides, were reported by
Ali et al.38,39
Actaea rubra: A
phytochemical fingerprint study by Jiang et al.36 found 12-14 as major phenolic acids in A. rubra. Depending on the material, the
triterpene glycosides identified were 1,
2, and 5-7, but 3 was notably
absent in all the samples analyzed.26,30,36,40 In addition, a number
of cimigenol derivatives and rubraside A have been isolated from the roots.40
According to one study, the compositions of A.
rubra and A. pachypoda are
similar and cannot be distinguished by chemical means.30
Actaea cimicifuga: A
large number of new triterpene glycosides have been reported from the roots and
rhizomes of A. cimicifuga, but
quantitative measurements of them are lacking. Only one paper presented a
quantitative analysis of the contents of the
major triterpene glycosides 2, 4, 5,
and 25-O-acetylcimigenol-3-O-xyloside.41 The same four
triterpene glycosides have also been reported by He et al,37 but 5 was absent in the material analyzed
by Wang et al.42 The main phenolic acids are 12 and 13, according to Jiang et al.36 The presence of 15 and 16, and the triterpene alkaloid
cimicifugadine (syn: cimicifine A), can be used to distinguish A. cimicifuga material from black
cohosh.23,34,37,42
Actaea dahurica:
Tang and Eisenbrand43 described 5,
7, cimigenol, dahurinol, shengmanol-3-O-xyloside, and 24-O-acetylanhydroshengmanol-3-O-xyloside from A. dahurica rhizomes. The presence of 2, 5, and 7 was reported by other research
groups.36,37 The main phenolic acids are 11, 12, and 14.36 As with A. cimicifuga, the presence of 15, 16, and cimicifugadine can be used to
distinguish A. dahurica material from
black cohosh.23,36,37,42
Actaea heracleifolia: Seven known triterpene glycosides – 5, 7, 8, 24-epi-24-O-acetylhydroshengmanol-3-O-xyloside, cimiaceroside B, 25-O-acetylcimigenol
3-O-β-d-xyloside, and 25-O-anhydrocimigenol
3-O-xyloside – were isolated from the rhizomes of A. heracleifolia.44 Importantly, both 1 and 2 reportedly do not occur in the species.23,36,37,42 The phenolic acids 12-14 have been described in A.
heracleifolia material by Jiang et al.36 at low levels, but not
by He et al.,37 who detected 2-feruloylpiscidic acid and
2-isoferuloylpiscidic acid instead. Since 15, 16, and cimicifugadine are found in A. heracleifolia roots and rhizomes,
these compounds can be used to distinguish it from black cohosh.23,36,42
Actaea simplex: Cycloartenol
triterpene glycosides are also predominant in the roots/rhizomes of A. simplex. Six new triterpenes have
been isolated by Kuang et al.45,46 The occurrence of 1, 2,
4, and 5 in the roots/rhizomes remains controversial,36,37,42 but
if confirmed, these compounds are present most likely at very low levels. The
phenolic acids 12-14 have been
reported from the roots and the rhizomes.36,37 The presence of 15 and 16, which have been reported from A. simplex, is indicative of adulteration of black cohosh with Chinese
Actaea species. However, the
identification of adulteration with A.
simplex should be based on a comparison of the overall chemical fingerprint
with authentic material.36,37
Acilepis aspera:
There are no published reports available in the chemical and other scientific
literature on the chemical composition of Acilepis
aspera roots.
Serratula
chinensis: The roots of S. chinensis were investigated by Ling
et al., who isolated seven ecdysteroids and five cembrosides, although only
three of them were subsequently identified.47,48 None of the
9,19-cycloartenol type triterpene glycosides or the phenolic acids typical for Actaea species have been reported from
this plant.
8.2 Laboratory Methods
Note: Unless otherwise
noted, all methods summarized below are based on using only the sub-aerial roots
and rhizomes of authentic black cohosh and its adulterants.
8.2.1 HPTLC
Methods from the following sources were evaluated in this
review: Upton,2 the USP 34,11 the EP 7.5,13
Gafner et al.,30 Wagner and Bladt,49 Zheng et al.,50
Ankli et al.,51 and Verbitski et al.52
Comments: The
HPTLC analysis of Actaea is a rare
instance where the majority of authors have relied on the same stationary and mobile
phases. The thorough validation, proven ability to detect adulteration, and
flexibility to target various phytochemicals depending on the detection
approach make the ethyl formate–toluene–formic acid (3:5:2, v/v) mobile phase
using HPTLC silica gel 60 F254 plates the
method of choice for HPTLC analysis of black cohosh (Figure 4).2,11,13,30,51-53
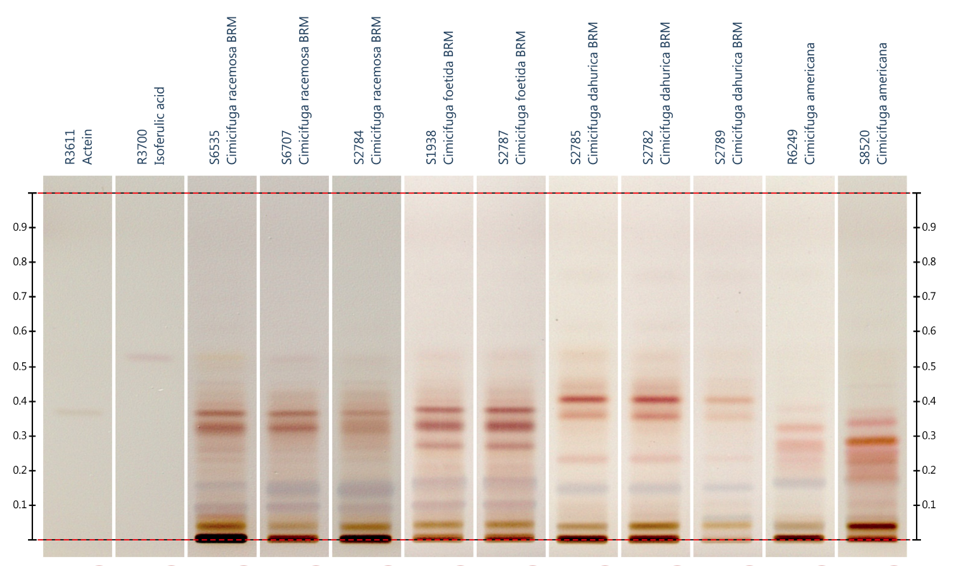
Figure
4:
HPTLC analysis of root extracts of black
cohosh, Actaea podocarpa (syn. Cimicifuga americana), and two Chinese Actaea spp., A. dahurica (syn. C. dahurica)
and A. cimicifuga (syn. C. foetida).53 Detection:
visible light after derivatization with sulfuric acid reagent. Image provided
by CAMAG (Muttenz, Switzerland).
Since method validations
were conducted using the sample preparation and detection system described in references
11 and 51, the consensus of authors and expert peer reviewers of this
Laboratory Guidance Document is that this procedure is the most suitable in a
routine QC lab. While this method is capable of distinguishing various Actaea species based on the chemical
fingerprint, the detection of adulterating species – in particular when such
species are added to A. racemosa – remains
challenging. Some of the related North American species exhibit a constituent profile similar to black cohosh; in
addition, the constituent profile may vary depending on the geographic location
and manufacturing process, although according to Eike Reich of CAMAG, the
chemical composition of black cohosh is rather consistent (Eike Reich e-mail
communication, November 19, 2014).
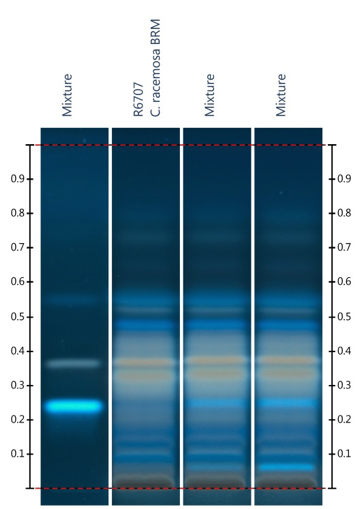
Figure
5:
HPTLC evaluation of black cohosh
adulteration with Actaea cimicifuga
and A. simplex. Detection: UV light
(366 nm) after derivatization with boric acid/oxalic acid reagent.53
Lane 1: cimifugin, actein, isoferulic acid (with increasing Rf value); lane 2: A. racemosa; lane 3: 5% A. cimicifuga in A. racemosa; lane 4: 5% A.
simplex in A. racemosa. Image
provided by CAMAG (Muttenz, Switzerland).
To detect adulteration with
Chinese Actaea species, the presence of
15 (found in, e.g., A. cimicifuga, A. dahurica, A. heracleifolia,
and A. simplex) can be verified using
boric acid-oxalic acid reagent reported by Ankli et al.51 The
application of boric acid-oxalic acid reagent leads to a strong fluorescence of
15 under UV light at 366 nm, and
allows the detection of as little as 1% of A.
cimicifuga and A. simplex in
black cohosh (Figure 5). For obvious
reasons, the boric acid-oxalic acid reagent does not allow the detection of
adulteration with Actaea species where
15 is absent (e.g., A. pachypoda, A. podocarpa, and A. rubra).
Detection of admixture of A.
heracleifolia or A. cimicifuga at
levels below 5% has been achieved using the antimony chloride detection
reagent.51
8.2.2
HPLC and UHPLC
Methods described in the following literature were
evaluated in this review: Upton,2 the USP 34,11 the EP 7.5,13 Ma et al.,23 Ganzera
et al.,25 Avula et al.,26,27 Li et al.,29
Gafner et al.,30 Jiang et al.,31,36,54 He et al.,37,55
Wang et al.,42 and Looney.56 Specific comments on
strengths and weaknesses of each of the methods are listed in Appendix 1, Table
4.
Comments: The
authentication and detection of black cohosh adulteration by HPLC or UHPLC should
be based on a chemical fingerprint, and the incoming raw material compared to
fingerprints from a number of representative authenticated samples by
statistical means. For routine quality control, a quick and easy sample
preparation method is provided in the European
Pharmacopoeia.13 The solvent of choice in most cases is a
mixture of MeOH-water (between 75 and 100% methanol, v/v) or EtOH-water (1:1 or
7:3, v/v).
Based on the run time, quality of separation, and extensive
validation, the HPLC-ELSD method presented in references 2 and 37 is a good
choice, but system suitability parameters have to be developed. Such parameters
have been specified only by the compendial methods outlined in the United States Pharmacopeia11
and the European Pharmacopoeia.13
If the run time is of essence, the conditions described by Avula et al.27
are by far the quickest, but require having a UHPLC instrument that can run
under higher pressure. However, no system suitability parameters are provided for
the method.
8.2.3
MS-Fingerprinting
Methods described in the following literature were
evaluated in this review: Huang et al.57 and Harnly et al.18
Comments:
Flow-injection mass spectrometry (FIMS), in which the ion spectrum is summed
over the injection interval, provides complex spectral fingerprints that, like
those for HPLC or NMR, can be used to compare unknown materials to a series of
authentic materials using statistical means. The sample preparation is identical
to that used for HPLC. Electrospray ionization was used by Huang et al.57
and Harnly et al.18 but the use of other ionization sources is
possible as well.
Normal mass calibration is required for alignment of the
spectral fingerprints. The authors used chemometric analysis, e.g., principal
component analysis (PCA), for visual inspection of the data and soft
independent modeling of class analogy (SIMCA) to provide statistical evaluation
of the degree of discrimination. As with other chemically based methods, this
approach works well with ingredients that are consistently processed in the
same way. This statistics-based authentication is state of the art for dietary
supplements. The analysis is very short and environmentally friendly due to low
solvent use. Data on method validation are not available. Initial costs for the
instrumentation are high.
8.2.4
NMR
One NMR-based authentication method was evaluated in this
review (Harnly et al.18).
Comments: Whole,
cut, or powdered samples are extracted using aqueous methanol followed by
drying and re-solvation in DMSO-d6.
For extracts, the material is directly dissolved in DMSO-d6. Harnly et al.18 use one-dimensional 1H-NMR
to establish spectral fingerprints of the crude extracts. The fingerprints allow
differentiating A. racemosa from
other Actaea species using statistical
evaluation, e.g., PCA and SIMCA, as well as identification and quantification
of some of the metabolites present in the samples.
The method provides
state-of-the-art statistics-based authentication. The results show that the NMR
approach is able to clearly distinguish A.
racemosa from other Actaea
samples. As with other statistics-based evaluations, added materials (e.g.,
carriers, processing aids) or variations in the manufacturing process will
modify the outcome of the PCA and thus may cluster the material outside the
acceptable range. Therefore, the construction of a library containing authenticated
materials of the same composition as the analyte is necessary. Expert analysts are
required to set up the right parameters and run the instrument. The analysis
time is short and ecologically responsible due to the low amount of solvent
used. As a result of the reproducibility using NMR, new samples can be directly
compared to samples run earlier without having to rerun the whole series.
System suitability for any botanical analysis is the same: the 1H
line shape and the 1H sensitivity have to comply with the probe
specifications. In addition, the temperature must be stable to 0.1°C. However,
the sample preparation for whole, cut, or powdered raw material is
time-consuming due to initial extraction time and the need to freeze-dry the
extract before analysis in order to avoid a large signal from residual water.
Data on method validation are not available. Initial costs for the
instrumentation are high.
9.
Conclusion
Authentication of cut or
powdered black cohosh rhizome is challenging due to the existence of closely
related and sometimes co-habiting Actaea
species with similar morphological features and chemical composition. On rare
occasions, phenotypes have displayed far greater variation than the genotypes.18
The need for sound analytical methods is further emphasized by the abundance of
materials from China sold as “black cohosh” but composed of root and sub-aerial
material from entirely different species. For authentication of raw material, a
combination of a physical assessment test (ideally using the whole plant)
and/or a genetic approach17,18 combined with chemical identification
methods is needed. For materials where DNA barcoding technology is applicable,
it has given the most accurate results. Rare cases of misidentification of A. racemosa, A. pachypoda, and A.
podocarpa rhizome based on chromatographic authentication methods have been
reported by Hartwig Sievers (e-mail communication, September 29, 2014) and
Harnly et al.18
Authentication or detection
of adulteration in extracts remains difficult since there is currently no method
available for a chemical compound or phytochemical class that is characteristic
for A. racemosa. (Some of the recently discovered alkaloids may be useful for
species authentication, but no methods have been published to date.) The
presence of 3 and absence of 15 and 16 are indicators for authentic black cohosh, but relying on the
presence/absence of a few marker compounds for the authentication of black
cohosh is insufficient. Any method for chemical authentication must be based on
a fingerprint, which means the entire spectrum of chemical compounds present
has to be evaluated using appropriate software for statistical analysis.
Several published HPTLC methods have shown their ability to distinguish black
cohosh and its major adulterants.2,11,13,30,51-53 The method of
choice for detection of adulterants added to authentic A. racemosa material by HPTLC is described by Ankli et al.51
Most of the published HPLC methods will be able to authenticate black cohosh, but
proving that it is only black cohosh (without any admixture of other material)
is a difficult task. When using HPLC, the sample preparation outlined in EP 7.513
combined with the chromatographic method reported in references 2 and 37 is a
good choice. Despite its main purpose for quantitative assessment, the
HPLC-ELSD method in the European
Pharmacopoeia13 has also been successfully applied to detect
adulteration of black cohosh and can be used for additional confirmation (Pilar
Pais personal communication, May 30, 2014). For increased specificity, the use
of an MS detector in addition to the ELSD should be considered.
Recent publications using
direct analysis by FIMS18,57 and NMR18 with subsequent
chemometric evaluation to distinguish between A. racemosa and other Actaea
species offer a unique approach in botanical authentication. These methods
combine simple sample preparation and rapid analysis. NMR offers unprecedented signal
stability (the intensity of the NMR signals change very little over time,
therefore allowing the comparison of spectra of new materials with archived
data, effectively eliminating the need to acquire data for standard materials
before each NMR experiment). MS and 1H-NMR have the proven ability
to fully characterize black cohosh and its adulterants. The results show that
FIMS and NMR (see below) perform equally well in distinguishing the various Actaea species; however, the results
also show the limitations of chemistry-based identification methods. Despite a
robust statistical evaluation of the results, one sample of A. pachypoda root clustered within the
black cohosh samples, and one sample of A.
racemosa root fell outside the 95% confidence interval set for
authentication of black cohosh. Both samples were correctly identified using a
DNA barcoding approach.18 These cases of misidentification may be
rare, and have been related to the unusual chemical composition of the
materials.
Note: A number of identity tests for black cohosh
materials are offered by third-party analytical laboratories. According to
input from six contract laboratories, the testing methods include microscopy,
DNA barcoding, HPTLC, and HPLC-UV. Additional testing methods (HPLC-MS or
near-infrared [NIR] methods) can be developed upon request.
Table
3. Comparison among the different approaches to authenticate A. racemosa.
|
Original name
|
Synonyms
|
New name
|
|
23-O-acetylshengmanol-3-O-xyloside
(7)
|
|
(23R)-23-acetoxy-(24S)-24,25-epoxy-(15R)-15-hydroxy-16-oxo-3-O-β-d-xylopyranosylactanoside
|
|
Actein (1)
|
Shengmating
|
(12R)-12-acetoxy-(24R,25S)-24,25-epoxy-(26R&S)-26-hydroxy-3-O-β-d-xylopyranosylacta-(16S,23R)-16,23;23,26-binoxoside
|
|
Cimigenol-3-O-arabinoside (6)
|
Cimiracemoside C,
cimicifugoside M
|
(15R)-15,25-dihydroxy-3-O-α-L-arabinopyranosylacta-(16S,23R,24S)-16,23;16,24-binoxoside
|
|
Cimigenol-3-O-xyloside (5)
|
Cimicifugol-xyloside, cimigenoside,
cimicifugoside
|
(15R)-15,25-dihydroxy-3-O-β-d-xylopyranosylacta-(16S,23R,24S)-16,23;16,24-binoxoside
|
|
Cimiracemoside A (3)
|
Cimiracemoside F
|
(12R)-12-acetoxy-7,8-didehydro-(23R,24R)-23,24-dihydroxy-3-O-β-d-xylopyranosylacta-(16S,22R)-16,23;22,25-binoxoside
|
|
23-Epi-26-deoxyactein (2)
|
26-Deoxyactein,
27-deoxyactein
|
(12R)-12-acetoxy-(24R,25R)-24,25-epoxy-3-O-β-d-xylopyranosylacta-(16S,23R)-16,23;23,26-binoxoside
|
aOnly
whole and cut and sifted (c/s) materials.
bIn particular, detection of adulteration in
mixtures of A. racemosa and related Actaea species, due to variability in
triterpene-glycoside composition within A.
racemosa populations and due to occurrence of same compounds in other Actaea species.
cCosts
for high-resolution mass spectrometers and NMR instruments are generally above US
$250,000. A low-cost 300 MHz NMR for natural products analysis can be obtained
for US $150,000.
10. References
1. McGuffin M, Kartesz JT, Leung AY, Tucker AO. Herbs of Commerce. 2nd ed. Silver
Spring, MD: American Herbal Products Association; 2000.
2. Upton R, ed. American Herbal Pharmacopoeia and
Therapeutic Compendium: Black Cohosh Rhizome Actaea racemosa L. syn. Cimicifuga racemosa (L.) Nutt.:
Standards of Analysis, Quality Control, and Therapeutics. Scotts
Valley, CA: American Herbal Pharmacopoeia; 2002. http://www.herbal-ahp.org/order_online.htm.
3.
Gardner ZE, Lueck L, Erhardt EB, Craker LE. A morphometric analysis of Actaea racemosa L. (Ranunculaceae). Journal of Medicinally Active Plants. 2012;1(2):47-59.
Available at: http://scholarworks.umass.edu/jmap/vol1/iss2/3/. Accessed October
26, 2015.
4. The Plant List. Version 1.1 (September 2013).
Available at: http://www.theplantlist.org.
Accessed October 26, 2015.
5. Tropicos.org. Missouri Botanical Garden. Available at:
http://www.tropicos.org.
Accessed October 26, 2015.
6. USDA PLANTS Database. Available at: http://plants.usda.gov. Accessed
October 26, 2015.
7. Pengelly A, Bennett K.
Appalachian plant monographs: Black cohosh Actaea racemosa L. Available
at: http://www.frostburg.edu/aces/appalachian-plants. Published
2012. Accessed October 26, 2015.
8. Cumberford G. EMI vs. EMA: “Economically motivated
integrity” vs. economically motivated adulteration in the natural products
supply chain. HerbalEGram. 2012;9(5).
9. Schulz V. Cimicifuga. In: Blaschek W, Ebel S, Hackenthal E, et al., eds. HagerROM 2006. Hagers Handbuch der Drogen
und Arzneistoffe. Berlin, Germany: Springer-Verlag GmbH; 2006.
10.
Hiller K. Cimicifugae racemosae rhizoma Traubensilberkerzen-Wurzelstock. In:
Wichtl M, ed. Teedrogen und Phytopharmaka.
4th ed. Stuttgart, Germany: Wissenschaftliche Verlagsgesellschaft mbH;
2002:660-661.
11. United States Pharmacopeial Convention. Black cohosh.
In: United States Pharmacopeia and
National Formulary (USP 34-NF 29). Rockville, MD: United States Pharmacopeial Convention.
2012:1083-1085.
12. Upton R, Graff A,
Jolliffe G, Länger R, Williamson E, eds. American Herbal Pharmacopoeia: Botanical
Pharmacognosy - Microscopic Characterization of Botanical Medicines. Boca Raton, FL: CRC Press; 2011:217-220.
13.
The European Directorate for the Quality of Medicines & HealthCare. European
Pharmacopoeia (EP 7.5). Black cohosh. Strasbourg, France:
Council of Europe; 2012:4552-4555.
14. Applequist WL. Rhizome and root anatomy of potential
contaminants of Actaea racemosa L.
(black cohosh). Flora. 2003;198(5):358-365. Abstract available
at: http://www.sciencedirect.com/science/article/pii/S036725300470078X.
Accessed October 26, 2015.
15. Xu
GJ, Xu LS, eds. Species Systematization and Quality Evaluation of Commonly
Used Chinese Traditional Drugs. South-China Edition, Volume I.
Fuzhou, China: Fujian Science and Technology Press; 1994.
16. Zerega NJC, Mori S, Lindqvist C, Zheng Q, Motley TJ.
Using amplified fragment length polymorphisms (AFLP) to identify black cohosh (Actaea racemosa). Econ Bot.
2002;56(2):154-164. Abstract available at: http://link.springer.com/article/10.1663%2F0013-0001(2002)056%5B0154%3AUAFLPA%5D2.0.CO%3B2. Accessed October 26,
2015.
17. Baker DA, Stevenson DW,
Little DP. DNA barcode identification of black cohosh herbal dietary supplements.
J AOAC Int. 2012;95(4):1023-1034.
Abstract available at: http://www.ncbi.nlm.nih.gov/pubmed/22970567.
Accessed October 26, 2015.
18. Harnly J, Chen P, Sun J, et
al. Comparison
of flow injection MS, NMR, and DNA sequencing: methods for identification and authentication
of black cohosh (Actaea racemosa). Planta Med. 2015; (in press).
19. Linde
H. Die
Inhaltsstoffe von Cimicifuga racemosa
4. Mitt.: Actein: Der Ring D und die Seitenkette. Arch
Pharm (Weinheim). 1968;301(2):120-138.
Abstract available at: http://onlinelibrary.wiley.com/doi/10.1002/ardp.19683010206/abstract.
Accessed October 26, 2015.
20. Linde H. Die Inhaltsstoffe von Cimicifuga racemosa. 5. Mitt.: 27-Desoxyacetylacteol. Arch Pharm (Weinheim). 1968;301(5):335-341.
Abstract available at: http://onlinelibrary.wiley.com/doi/10.1002/ardp.19683010504/abstract. Accessed October 26, 2015.
21. Struck D, Tegtmeier M, Harnischfeger G. Flavones in
extracts of Cimicifuga racemosa. Planta Med. 1997;63(3):289.
22.
Kruse SO, Löhning A, Pauli GF, Winterhoff H, Nahrstedt A. Fukiic and piscidic
acid esters from the rhizome of Cimicifuga
racemosa and the in vitro estrogenic activity of fukinolic acid. Planta Med. 1999;65(8):763-764. Abstract
available at: https://www.thieme-connect.com/DOI/DOI?10.1055/s-2006-960862. Accessed October
26, 2015.
23. Ma C, Kavalier AR, Jiang B, Kennelly EJ. Metabolic profiling of Actaea (Cimicifuga)
species extracts using high performance liquid chromatography coupled with
electrospray ionization time-of-flight mass spectrometry. J Chromatogr A. 2011;1218(11):1461-1476. Available at:
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3389552/pdf/nihms275630.pdf.
Accessed October 26, 2015.
24. Nikolić D, Gödecke T, Chen
S-N, et al. Mass spectrometric dereplication of nitrogen-containing
constituents of black cohosh (Cimicifuga racemosa L.). Fitoterapia. 2012;83(3):441-460. Available at: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3341503/pdf/nihms343193.pdf. Accessed October 26, 2015.
25. Ganzera M, Bedir E, Khan IA. Separation of Cimicifuga
racemosa triterpene glycosides by reversed phase high performance liquid
chromatography and evaporative light scattering detection. Chromatographia. 2000;52(5-6):301-304.
Abstract available at: http://link.springer.com/article/10.1007/BF02491021. Accessed October 26, 2015.
26. Avula B, Ali Z, Khan IA. Chemical fingerprinting of Actaea racemosa
(black cohosh) and its comparison study with closely related Actaea
species (A.
pachypoda,
A.
podocarpa,
A.
rubra)
by HPLC. Chromatographia.
2007;66(9-10):757-762. Abstract available at: http://link.springer.com/article/10.1365%2Fs10337-007-0384-6. Accessed October 26, 2015.
27. Avula B, Wang Y-H,
Smillie TJ, Khan IA. Quantitative determination of triterpenoids and formononetin
in rhizomes of black cohosh (Actaea
racemosa) and dietary supplements by using UPLC-UV/ELS detection and
identification by UPLC-MS. Planta Med.
2009;75(4):381-386. Abstract available at: https://www.thieme-connect.com/DOI/DOI?10.1055/s-0028-1088384.
Accessed October 26, 2015.
28. Shao Y, Harris A, Wang M, et al. Triterpene
glycosides from Cimicifuga racemosa. J Nat Prod. 2000;63(7):905-910. Abstract
available at: http://pubs.acs.org/doi/abs/10.1021/np000047y.
Accessed October 26, 2015.
29. Li W, Chen S, Fabricant D, et al. High-performance liquid
chromatographic analysis of black cohosh (Cimicifuga racemosa)
constituents with in-line evaporative light scattering and photodiode array
detection. Anal Chim Acta.
2002;471(1):61-75. Abstract available at: http://www.sciencedirect.com/science/article/pii/S0003267002007766.
Accessed October 26, 2015.
30. Gafner S, Sudberg S, Sudberg ÉM, Villinski
JR, Gauthier R, Bergeron C. Chromatographic fingerprinting as a means of
quality control: Distinction between Actaea racemosa and four different Actaea species. Acta
Hortic. 2006;720:83-94.
Abstract available at: http://www.actahort.org/books/720/720_8.htm.
Accessed October 26, 2015.
31. Jiang
B, Kronenberg F, Nuntanakorn P, Qiu M-H, Kennelly EJ. Evaluation of the botanical authenticity and
phytochemical profile of black cohosh products by high-performance liquid
chromatography with selected ion monitoring liquid chromatography-mass spectrometry. J Agric Food Chem. 2006;54(9):3242-3253. Available at: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3204370/pdf/nihms23156.pdf. Accessed October 26, 2015.
32. Cicek SS, Aberham A, Ganzera M, Stuppner H.
Quantitative analysis of cycloartane glycosides in black cohosh rhizomes and dietary supplements
by RRLC-ELSD and RRLC-qTOF-MS. Anal
Bioanal Chem. 2011;400(8):2597-2605. Abstract available at: http://link.springer.com/article/10.1007%2Fs00216-010-4068-y.
Accessed October 26, 2015.
33. European Scientific
Cooperative on Phytotherapy (ESCOP) Monographs. Cimicifugae Rhizoma: Black Cohosh. Exeter, United Kingdom: ESCOP
Secretariat; 2011.
34. Nuntanakorn P, Jiang B, Yang H, Cervantes-Cervantes M, Kronenberg F, Kennelly EJ. Analysis of polyphenolic
compounds and radical scavenging activity of four American Actaea species. Phytochem Anal.
2007;18(3):219-228. Available at: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2981772/pdf/nihms-23157.pdf.
Accessed October 26, 2015.
35. Qiu F, McAlpine JB, Krause EC, Chen S-N, Pauli GF. Pharmacognosy
of black cohosh: the phytochemical and biological profile of a major botanical
dietary supplement. In: Kinghorn AD, Falk H, Kobayashi J, eds. Progress
in the Chemistry of Organic Natural Products, Vol. 99. Cham, Switzerland: Springer International
Publishing; 2014:1-68.
36. Jiang B, Ma C,
Motley T, Kronenberg F, Kennelly EJ. Phytochemical fingerprinting to thwart
black cohosh adulteration: a 15 Actaea species
analysis. Phytochem Anal. 2011;22(4):339-351.
Abstract available at: http://onlinelibrary.wiley.com/doi/10.1002/pca.1285/abstract. Accessed October 26, 2015.
37. He K, Pauli GF, Zheng B, et al. Cimicifuga species identification by high performance liquid
chromatography–photodiode array/mass spectrometric/evaporative light scattering
detection for quality control of black cohosh products. J Chromatogr A. 2006;1112(1-2):241-254. Available at: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1847404/pdf/nihms14941.pdf. Accessed October 26, 2015.
38. Ali Z, Khan SI,
Fronczek FR, Khan IA. 9,10-seco-9,19-Cyclolanostane arabinosides from the roots of Actaea podocarpa. Phytochemistry.
2007;68(3):373-382. Abstract available at: http://www.sciencedirect.com/science/article/pii/S0031942206006637. Accessed October 26, 2015.
39. Ali
Z, Khan SI, Khan
IA. New cycloartane-type triterpene arabinosides from the roots of Actaea
podocarpa and their biological study. Planta Med. 2007;73(7):699-703. Abstract available at: https://www.thieme-connect.com/DOI/DOI?10.1055/s-2007-981533. Accessed October 26, 2015.
40. Ali
Z, Khan SI, Khan
IA. Phytochemical study of Actaea rubra and biological screenings
of isolates. Planta Med. 2006;72(14):1350-1352. Abstract available at: https://www.thieme-connect.com/DOI/DOI?10.1055/s-2006-951696. Accessed October 26, 2015.
41. Gai Y-Y, Liu W-H, Sha C-J, et al. Pharmacokinetics and
bioavailability of cimicifugosides after oral administration of Cimicifuga
foetida L. extract to rats. J
Ethnopharmacol. 2012;143(1):249-255. Abstract available at: http://www.sciencedirect.com/science/article/pii/S037887411200431X. Accessed October 26, 2015.
42.
Wang H-K, Sakurai N, Shih CY, Lee K-H. LC/TIS-MS fingerprint profiling of Cimicifuga species and analysis of 23-epi-26-deoxyactein
in Cimicifuga racemosa commercial
products. J Agric Food Chem.
2005;53(5):1379-1386. Abstract available at: http://pubs.acs.org/doi/abs/10.1021/jf048300d. Accessed October 26, 2015.
43. Tang W, Eisenbrand G. Chinese Drugs of Plant Origin: Chemistry, Pharmacology, and Use in
Traditional and Modern Medicine. Berlin, Germany: Springer-Verlag; 1992:315-317.
44. Lee JH, Cuong TD,
Kwack SJ, et al. Cycloartane-type
triterpene glycosides from the rhizomes
of Cimicifuga
heracleifolia and their anticomplementary activity. Planta Med. 2012;78(12):1391-1394. Abstract available
at: https://www.thieme-connect.com/DOI/DOI?10.1055/s-0032-1314980. Accessed October 26, 2015.
45. Kuang H, Su Y, Yang B, et al. Three new cycloartenol triterpenoid saponins from the roots of Cimicifuga simplex
Wormsk. Molecules.
2011;16(6):4348-4357. Available at: http://www.mdpi.com/1420-3049/16/6/4348. Accessed October 26, 2015.
46. Kuang H, Su Y, Wang Q, et al. Three new cycloartenol
glycosides from the roots of Cimicifuga simplex. Planta Med. 2012;78(6):622-625.
Abstract available at: https://www.thieme-connect.com/DOI/DOI?10.1055/s-0031-1298224. Accessed October 26, 2015.
47. Ling
T-J, Ma W-Z, Wei X-Y. Ecdysteroids from the roots of Serratula chinensis.
J Trop Subtrop Botany. 2003;11(2):143-147. Available at: http://jtsb.scib.ac.cn/jtsb_en/ch/reader/view_abstract.aspx?flag=1&file_no=200302010&journal_id=jtsb_cn. Accessed October
26, 2015.
48. Ling T, Xia T, Wan X, Li D, Wei X. Cerebrosides from the roots of Serratula
chinensis. Molecules. 2006;11(9):677-683. Available at: http://www.mdpi.com/1420-3049/11/9/677. Accessed October
26, 2015.
49.
Wagner H, Bladt S. Plant Drug Analysis: A
Thin Layer Chromatography Atlas. 2nd ed. Berlin, Germany: Springer-Verlag;
1996:335-339.
50. Zheng QY, He K, Pilkington L, Shao Y, Zheng B. CimiPure (Cimicifuga racemosa): A standardized black cohosh extract with
novel triterpene glycoside for menopausal women. In: Shahidi F, Ho C-T, eds. Phytochemicals
and Phytopharmaceuticals. Champaign, IL: AOCS Press; 2000:368.
51. Ankli
A, Reich E, Steiner M. Rapid high-performance thin-layer chromatographic method
for detection of 5% adulteration of black cohosh with Cimicifuga foetida, C.
heracleifolia, C. dahurica, or C. americana. J
AOAC Int. 2008;91(6):1257-1264.
Abstract available at: http://www.ncbi.nlm.nih.gov/pubmed/19202784.
Accessed October 26, 2015.
52. Verbitski
SM, Gourdin GT, Ikenouye LM, McChesney JD, Hildreth J. Detection of Actaea racemosa adulteration by
thin-layer chromatography and combined thin-layer
chromatography-bioluminescence. J AOAC Int. 2008;91(2):268-275. Available at: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4115334/pdf/nihms66104.pdf.
Accessed October 26, 2015.
53.
International Association for the Advancement of High Performance Thin Layer
Chromatography. Methods: Cimicifuga
racemosa (black cohosh), identification – rhizome. HPTLC Association
website. Available at: http://www.hptlc-association.org/methods.cfm. Published
July 21, 2014. Accessed October 26, 2015.
54. Jiang
B, Yang H, Nuntanakorn P, Balick MJ, Kronenberg F, Kennelly EJ. The value of
plant collections in ethnopharmacology: a case study of an 85-year-old black
cohosh (Actaea racemosa L.) sample. J
Ethnopharmacol. 2005;96(3):521-528.
Abstract available at: http://www.sciencedirect.com/science/article/pii/S0378874104005045.
Accessed October 26, 2015.
55. He K, Zheng B, Kim CH, Rogers L, Zheng Q. Direct analysis
and identification of triterpene glycosides by LC/MS in black cohosh, Cimicifuga racemosa,
and in several commercially available black cohosh products. Planta Med. 2000;66(7):635-640. Abstract
available at: https://www.thieme-connect.com/DOI/DOI?10.1055/s-2000-8619. Accessed October 26, 2015.
56. Looney
PM. Quantification and characterization of biologically active components of Actaea racemosa L. (black cohosh) for
identifying desirable plants for cultivation. PhD thesis. Cullowhee, NC:
Western Carolina University; 2012. Available at: http://libres.uncg.edu/ir/wcu/f/Looney2012.pdf. Accessed
October 26, 2015.
57. Huang H, Sun J, McCoy J-A, et al. Use of flow injection mass
spectrometric fingerprinting and chemometrics for differentiation of three
black cohosh species. Spectrochim Acta Part
B At Spectrosc. 2015;105:121-129. Abstract available at: http://www.sciencedirect.com/science/article/pii/S0584854714002754. Accessed
October 26, 2015.
58. LaBudde RA, Harnly JM. Probability of identification: a statistical
model for the validation of qualitative botanical identification methods. J AOAC Int. 2012;95(1):273-285.
Available at: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3620024/pdf/nihms450355.pdf. Accessed
October 26, 2015.
Appendix
1
Table 4: Comments on the
published HPLC methods for A. racemosa.
|
Reference
|
Comments
|
|
Upton,2 He37
|
This is
a validated HPLC-ELSD method with an acceptable duration of run and good peak
shapes. The fingerprint data show an A.
racemosa triterpene-glycoside pattern that is distinct from 2 North
American and 7 Asian Actaea
species. The sample preparation is lengthy based on the 24-hr extraction
period, but consists only of a few handling steps. In order to extend column
life, a mobile phase containing 0.1% formic acid is preferred. There is no information
on peak identity using the validated ELSD detection and not all peaks are
well separated.
|
|
USP,11 Li,29 Gafner30
|
This
HPLC-ELSD method has good peak shapes and a reasonable separation. It has
been adopted as official method by the USP. The USP monograph contains detailed parameters for system suitability. However,
the sample preparation is labor-intensive and the HPLC run time of 70 min is unnecessarily
long since no peaks elute after 55 min. Using the validated ELSD detection
will not provide information on peak identity. Not all peaks are well
separated.
|
|
EP13
|
This HPLC-ELSD method has
been validated and contains detailed parameters for system suitability. The sample
preparation is quick and easy. The run time is reasonably short. The method was
developed to quantify triterpene glycosides and there is no published data on
its ability to authenticate black cohosh (the EP 7.5 standard method for
authentication relies on HPTLC). Using the validated ELSD detection will
not provide information on peak identity.
|
|
Ma23
|
This HPLC-MS method has been tested on 9 A. racemosa populations and 15
additional Actaea species. The
statistical evaluation of HPLC-TOFMS fingerprints using PCA is state of the
art. The results show that authentication of A. racemosa is possible based on the presence of 6 combined with the absence of 15
and 16. Sample preparation
is time-consuming and labor-intensive. An HPLC system that can handle ternary
solvent systems is required. The separation is reasonable, but the run time
of 103 min is quite long. The method has not
been validated.
|
|
Ganzera25
|
This
HPLC-ELSD method has a quick and simple sample preparation procedure. The run
time is reasonably short but the chromatogram shows some overlapping peaks. The
method has proven its ability to differentiate A. racemosa from A.
dahurica and A. cimicifuga, but
the data are limited to one sample per species. The composition of the mobile
phase requires an HPLC system that can handle ternary solvent systems. The use
of an ELSD detector will not provide information on peak identity. The method
has not been validated.
|
|
Avula26
|
This is
another HPLC-ELSD approach with a quick sample preparation method. The conditions
lead to a good separation for 1, 2, and 3, but the chromatogram shows some overlapping of later eluting
compounds, and the run time is long (75 min). The method has been validated
(according to the authors) for authentication by testing 4 populations of A. racemosa and 3 North American Actaea species, but not in terms of
quantitative analysis. The composition of the mobile phase requires an HPLC
system that can handle ternary solvent systems. The use of an ELSD detector
will not provide information on peak identity.
|
|
Avula27
|
This is
a validated method combining UHPLC with three detectors (UV/Vis, ELSD, and
MS). The chromatogram shows good peak shapes and separation for 1, 2, and 3 (the
chromatograms look simple compared to those in reference 23 or 29). The run
time is short and the sample preparation method is quick. The method has been
shown to distinguish A. racemosa
and 3 related North American Actaea
species.
|
|
Jiang,31 Jiang,54
He55
|
The
HPLC-UV (detection at 203 nm) method has been validated,31 despite
some unresolved peaks. It has proven the ability to identify adulteration if
HPLC-MS (triterpene glycosides) and HPLC-UV methods (phenolic acids) are used
in combination. However, the sample preparation technique using chloroform is
labor-intensive and the HPLC run time lengthy (65 min).
|
|
Jiang36
|
The HPLC-UV (detection at 203 nm) method has been
validated, despite some unresolved peaks. The duration of the HPLC run is
acceptable. The method is capable of identifying adulteration based on the fingerprint
analysis, presence of 3, and
absence of 15 (however, A. pachypoda also contains 3 and lacks 15). The sample preparation technique using chloroform is
labor-intensive.
|
|
Wang,42
fingerprint
|
The
authors present an HPLC-MS fingerprinting method that is able to distinguish
between A. racemosa and 6 Asian Actaea species. The analysis of 6
commercial samples with rather different triterpene-glycoside fingerprints
shows the challenges of correct authentication. The chromatographic system
leads to good peak shapes and a reasonable separation, but has a long run
time of 93 min. The sample preparation is labor-intensive and uses chloroform.
The injection volume is high (50 µL of sample in methanol) for initial
conditions consisting of MeCN-10 mM
ammonium acetate in water (5:95, v/v). Crucial information on MS
parameters is lacking and the method has not been validated.
|
|
Looney56
|
The
author analyzed a large number (20) of A.
racemosa populations with this HPLC-ELSD method. The sample preparation is
easy but long. The run time and separation are reasonable, although there are
some overlapping peaks. The use of an ELSD detector will not provide
information on peak identity. The method has
not proven its ability to detect adulteration with other species and has
not been completely validated.
|